Frame Selection
It may be desirable not to save all acquired frames to disk. Perhaps certain imaging channels may be relevant only for certain acquisition volumes and ignored for others. Or perhaps the discard frames from a FastZ acquisition is truly not useful. The File Selection feature may be used in these cases where more advanced composition of saved frames may be important, either simply to reduce file footprints on disk, or to provide a basic demultiplexed subset of frame data that may be saved in one or more output files.
Warning
This feature only exists for NI/VDAQ Resonant Scanning modes (Resscan, and RGGScan respectively). Frame selection in Linear Scanning or Line Scanning modes are not currently supported.
Prerequisites and Concepts
Frame selection is only enabled if logging is enabled. When frame logging is enabled, the Frame Selection feature is ignored.
Suffixes
Frame selections partition a stream of frames into multiple files by modifying the file name. These are called file suffixes and serve to identify which file the each frame is being selected to write to. Each file suffix is independent of the other and cannot share channel or frame selections.
File suffixes are assigned in the NI ResScan or VDAQ RGGScan Acquisition object under the FrameSelectFileSuffixList property. Multiple file suffix groups should be specified as a cell array of characters.
Selection Groups
Selecting individual frames per acquisition is memory-inefficient, especially when doing so with a large number of frames. To reduce this inefficiency, frames are selected using selection group which consists of an array of structs with the following format:
struct( ...
StartIndex = {2, 1} ...
, EndIndex = {Inf, Inf} ...
, StrideIndex = {2, 2})
The
StartIndexfield indicates the frame index to start the selection group.The
EndIndexfield indicates the ending index. As in the example above, replacing the value withInfis a valid shorthand for indicating that the selection group extends to the end of the acquisition.The
StrideIndexfield indicates the number of output frames to skip between the start index and the end index.
The number of struct groups must match the number of file suffixes. The field values may be scalar or vectors indicating a union of selection groups for, say, special cases where a single set of start, end, or stride indices. In the case where the selection group is represented by vectors, the number of elements in the vector must match across start, end, and stride indices. Selection groups are assigned to the FrameSelectSelectionList property in the NI or VDAQ resonant scanner acquisition objects. For both ResScan (NI) and RGGScan (VDAQ) the fields will be found here: hSI.hScan2D.hAcq.FrameSelectSelectionList.
Averaging
Frame selections operate on indices after frames are averaged. Pre-averaged frames cannot be selected using the frame selection feature. This may mean that frame index of 3 may not be the exact same acquisition frame if frame averaging is enabled. For assistance in visualizing actual selected frames, see Using the Graphical Viewer.
Channels
Channels can also be selected for frame selection. This feature is orthogonal to the frame selection feature though it operates using a similar syntax.
The FrameSelectChannelIndexList field is a cell array of indices indicating all currently saved channels that should be saved to this file suffix. This field is required and the number of elements within the cell array must equal the number of file suffixes.
Warning
If the channel indices are not specified, no frame data may be saved to file. The exact channel indices must be indicated in order for the frame selection group to properly save the right channels to the file.
Scripting Overview
Editing Selection Groups
The following code adds a file suffix labelled wave_900nm which selects frames 1-4 and all odd frames afterwards. The selection group also select channels 1 and 2 for saving to the file with the suffix wave_900nm.
AcquisitionObject = hSI.hScan2D.hAcq;
AcquisitionObject.FrameSelectFileSuffixList = {'wave_900nm'};
AcquisitionObject.FrameSelectChannelIndexList = {[1,2]};
AcquisitionObject.FrameSelectSelectionList = struct('StartIndex', [1, 3], 'EndIndex', [3, Inf], 'StrideIndex', [1, 2]);
Starting a logging acquisition after running the above script will now generate a file of the form <filestem>_wave_9000nm_00001.tif which only selects the configured groups.
Using the Graphical Viewer
To aide in visualizing the effects of user selections, an interactive graphical interface has been provided which helps indicate which frames of an acquisition will be logged to which file. The following command in the Command Window will open the frame selection graphical viewer:
scanimage.guis.openFrameSelectionWindow();
Warning
The graphical viewer window requires that ScanImage is fully open and that the currently selected scanner supports Frame Selections.
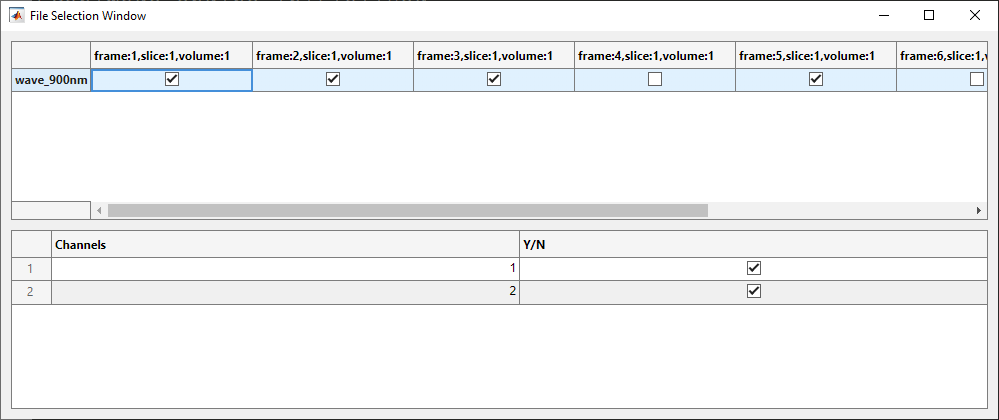
Graphical Viewer Sample
A sample of the frame selection graphical viewer. The settings should match the code from the Editing Selection Groups section